3.10 Detecting Outliers
The PCA above uses only the top 500 most variable genes (DESeq2 default). Here we run PCA on the full VST matrix and inspect a scree plot and biplot to assess whether any single sample drives an unusual amount of variance, a common sign of a technical outlier.
pca_full <- prcomp(t(assay(vsd)))
fviz_screeplot(pca_full, addlabels = TRUE,
main = "Scree plot — variance per PC")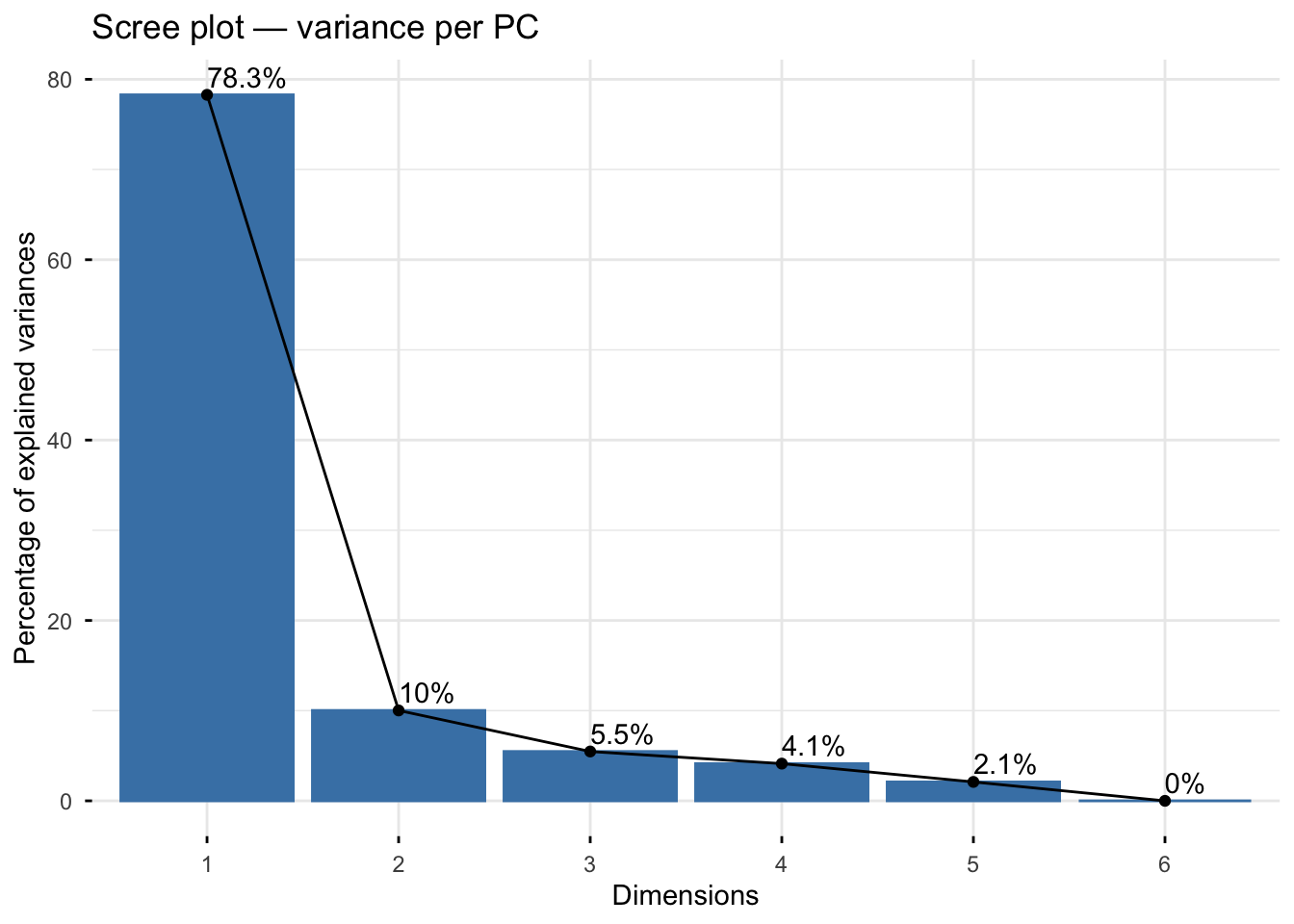
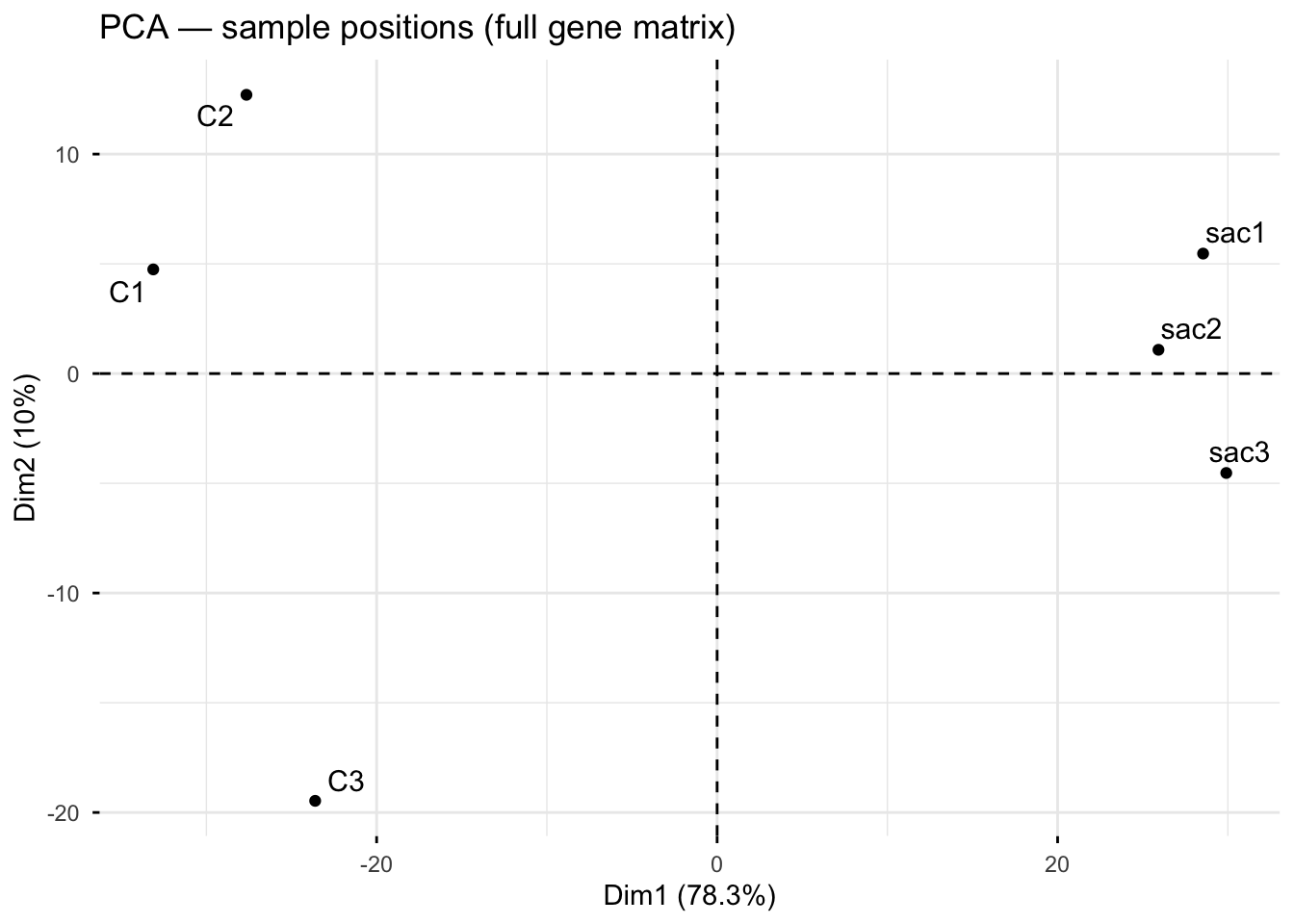
fviz_pca_ind(pca_full, geom = c("point", "text"), repel = TRUE,
title = "PCA — sample positions (full gene matrix)")