5.4 Part 1 — Over-Representation Analysis (ORA)
What is ORA?
ORA asks: “Are my significant DE genes enriched in any pathway more than expected by chance?”
It uses a hypergeometric test (equivalent to a one-sided Fisher’s exact test):
- Query set — significant DE genes (padj < 0.05, |LFC| >= 1), converted to gene symbols
- Background — all genes tested by DESeq2, converted to gene symbols
- Gene sets — KEGG pathways
- FDR correction — empirical FDR (eFDR) via resampling, which accounts for interdependence between gene sets
Key assumption: ORA treats all significant genes equally — it ignores magnitude or direction of fold change. This is why we also run GSEA (Part 2), which uses the full ranked list.
5.4.1 Prepare Input
sig_genes <- res_sig_sym$gene
all_genes <- res_df_sym$gene
cat("Query set (significant genes) :", length(sig_genes), "\n")## Query set (significant genes) : 646## Pool (all tested genes) : 3698cat("Overlap with first pathway :",
length(intersect(sig_genes, kegg_gmt$list_of_values[[1]])), "genes\n")## Overlap with first pathway : 7 genesTip: Always use all tested genes as the background, not the full
genome. DESeq2 already filtered to expressed genes, so res_df_sym is the
correct pool. Using the full genome inflates the background and produces over-optimistic
p-values.
5.4.2 Run ORA
set.seed(42)
ora_model <- ora(
gmt = kegg_gmt,
element_names = sig_genes,
background_element_names = all_genes,
p_value_adjustment_method = "eFDR",
number_of_permutations = 1000
)
ora_results <- mulea::run_test(ora_model)
cat("Pathways tested :", nrow(ora_results), "\n")## Pathways tested : 121## Significant (eFDR < 0.05): 125.4.3 Filter and Inspect Significant Pathways
ora_sig <- ora_results %>%
filter(eFDR < 0.05) %>%
arrange(eFDR)
cat("Significant ORA pathways:", nrow(ora_sig), "\n")## Significant ORA pathways: 12DT::datatable(
ora_sig %>%
select(ontology_id, ontology_name, nr_common_with_tested_elements,
nr_common_with_background_elements, p_value, eFDR) %>%
mutate(across(where(is.numeric), \(x) round(x, 4))),
rownames = FALSE,
colnames = c("KEGG ID", "Pathway", "Hits in query",
"Hits in background", "p-value", "eFDR"),
extensions = c("Buttons", "Scroller"),
options = list(
dom = "Bfrtip",
buttons = c("copy", "csv"),
scrollX = TRUE,
scrollY = 300,
scroller = TRUE
),
caption = "Significant ORA pathways — treatment vs control (eFDR < 0.05)"
)Visualise ORA Results
5.4.4 Bar Plot — Enriched Pathways
ora_bar <- ora_sig %>%
slice_min(eFDR, n = 20) %>%
mutate(ontology_name = fct_reorder(ontology_name, -log10(eFDR)))
ora_barplot <- ggplot(ora_bar,
aes(x = -log10(eFDR),
y = ontology_name,
fill = -log10(eFDR))) +
geom_col(width = 0.7) +
geom_vline(xintercept = -log10(0.05), linetype = "dashed",
color = "black", linewidth = 0.5) +
scale_fill_gradient(low = "#92C5DE", high = "#CA0020") +
theme_bw() +
theme(legend.position = "none") +
labs(
title = "ORA — Enriched KEGG Pathways (eFDR < 0.05)",
subtitle = "Treatment vs Control | E. coli MG1655",
x = "-log10(eFDR)",
y = NULL
)
ora_barplot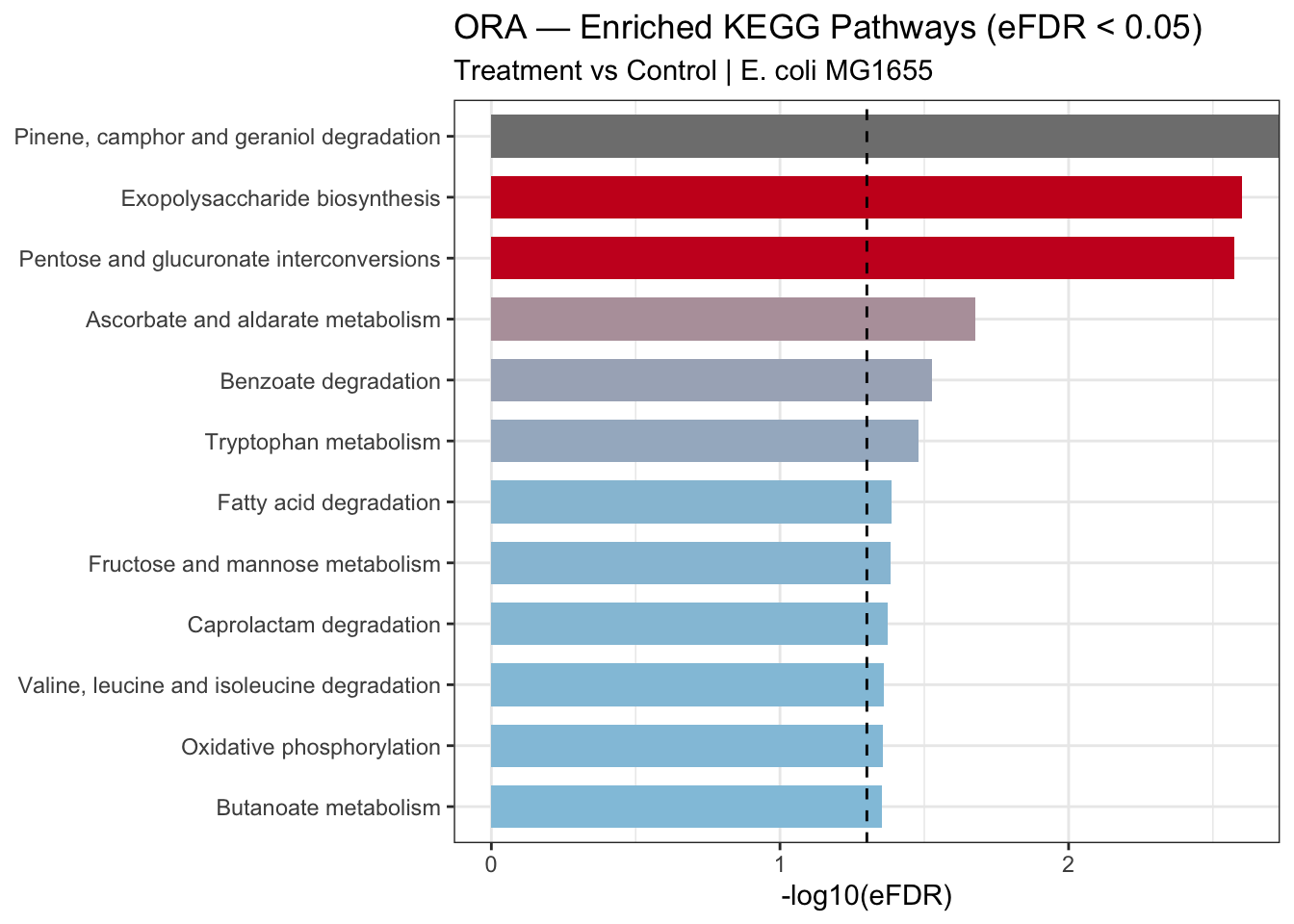
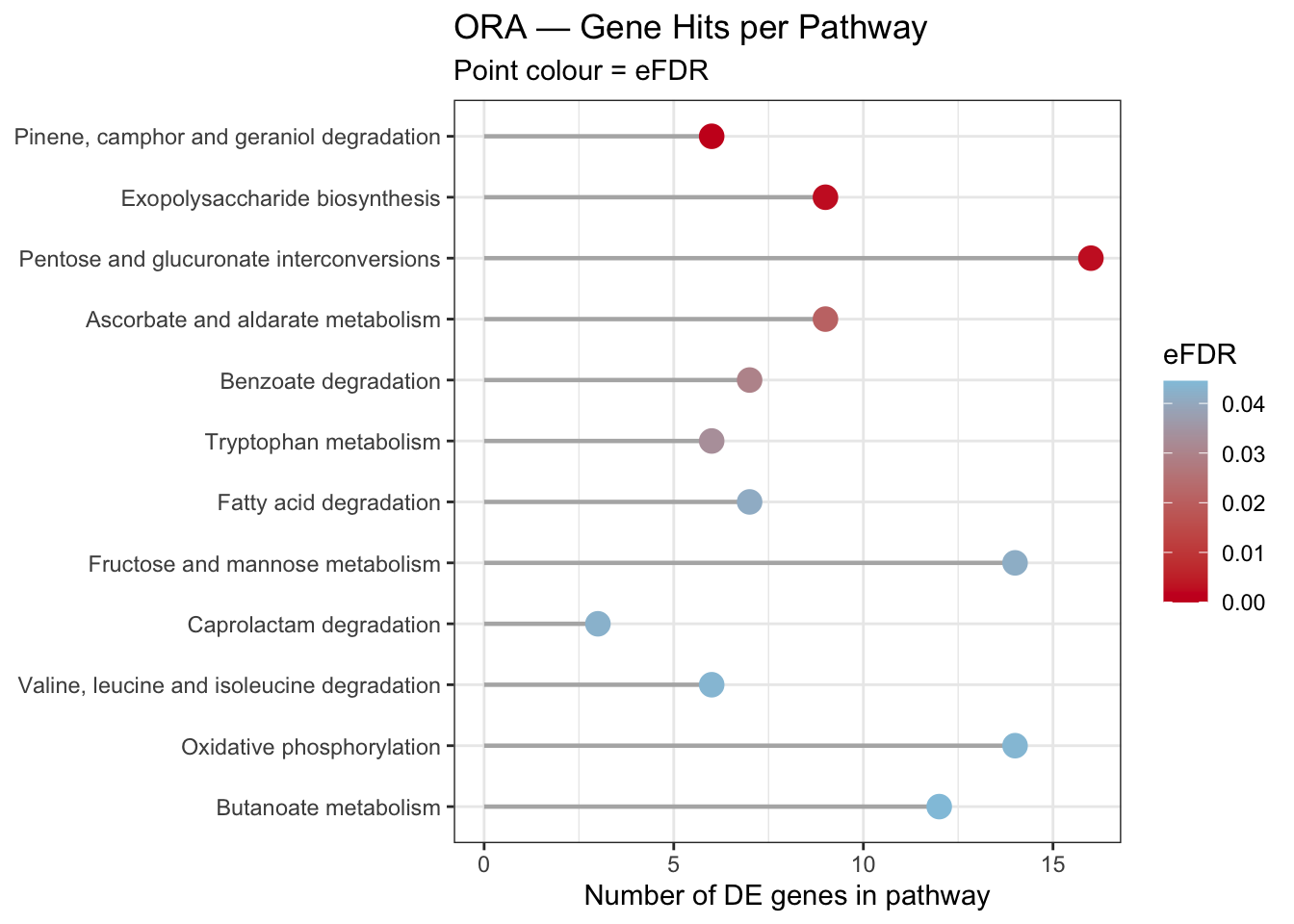
5.4.5 Lollipop Plot — Gene Hits per Pathway
ora_lollipop <- ggplot(ora_bar,
aes(x = nr_common_with_tested_elements,
y = ontology_name,
color = eFDR)) +
geom_segment(aes(x = 0,
xend = nr_common_with_tested_elements,
yend = ontology_name),
linewidth = 0.8, color = "grey70") +
geom_point(size = 4) +
scale_color_gradient(low = "#CA0020", high = "#92C5DE", name = "eFDR") +
theme_bw() +
labs(
title = "ORA — Gene Hits per Pathway",
subtitle = "Point colour = eFDR",
x = "Number of DE genes in pathway",
y = NULL
)
ora_lollipop