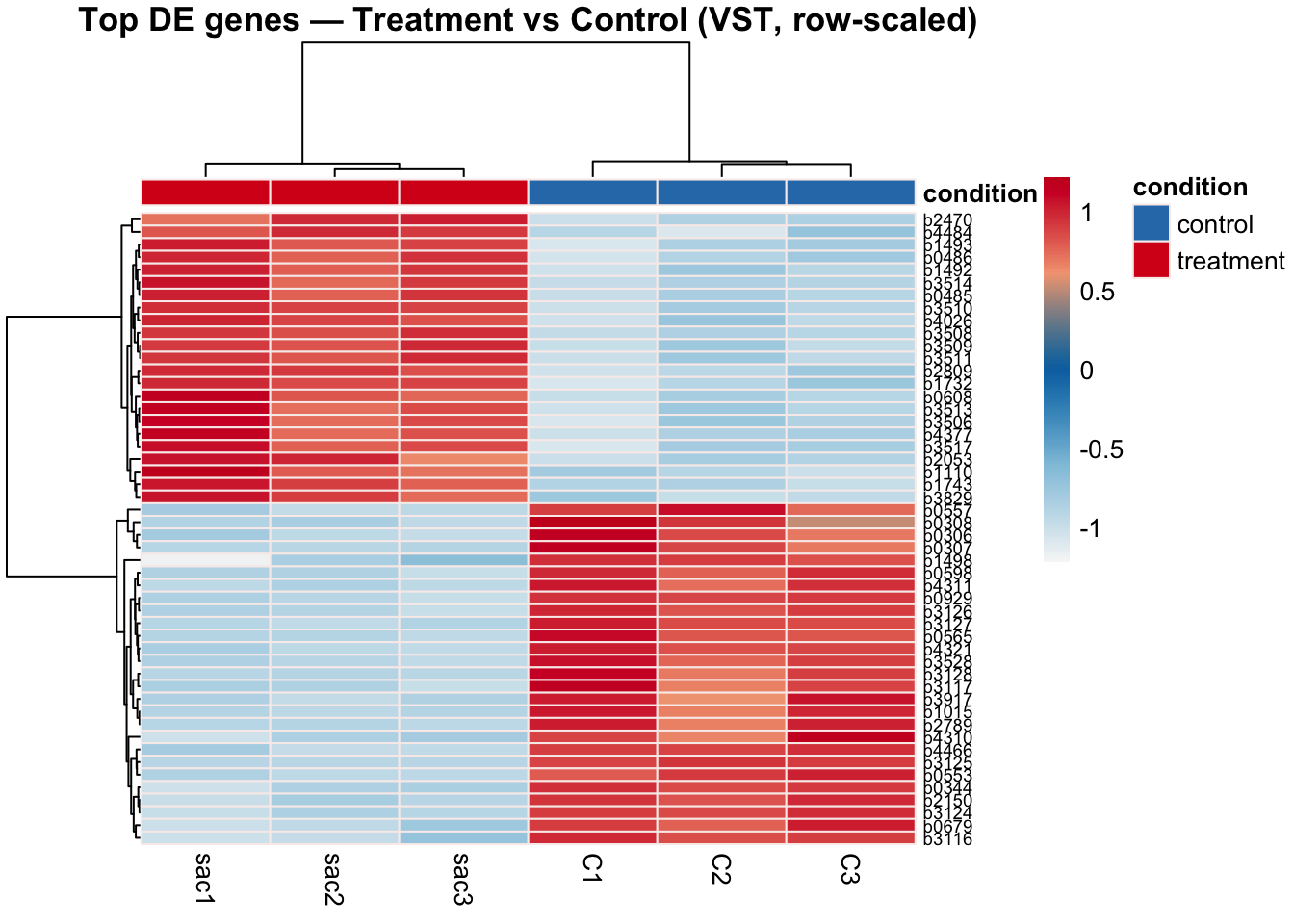
4.8 Heatmap of Significant Genes
Clustering significant DE genes across samples provides a gene-level view of the expression differences. Genes should show clear block structure: high expression in treatment and low in control (or vice versa). Replicates within each condition should cluster together.
vsd <- varianceStabilizingTransformation(dds, blind = FALSE)
sig_genes <- res_sig %>%
slice_min(padj, n = min(50, nrow(res_sig))) %>%
pull(gene)
df_anno <- as.data.frame(colData(vsd)[, "condition", drop = FALSE])
anno_colors <- list(condition = cols_condition)
colsHeat <- c("#F7F7F7", "#92C5DE", "#0571B0", "#F4A582", "#CA0020")
pheatmap(
assay(vsd)[sig_genes, ],
cluster_cols = TRUE,
cluster_rows = TRUE,
scale = "row",
clustering_distance_rows = "euclidean",
clustering_distance_cols = "euclidean",
annotation_col = df_anno,
annotation_colors = anno_colors,
show_colnames = TRUE,
show_rownames = TRUE,
color = colorRampPalette(colsHeat)(255),
border_color = "#f8edeb",
fontsize_row = 7,
main = "Top DE genes — Treatment vs Control (VST, row-scaled)"
)