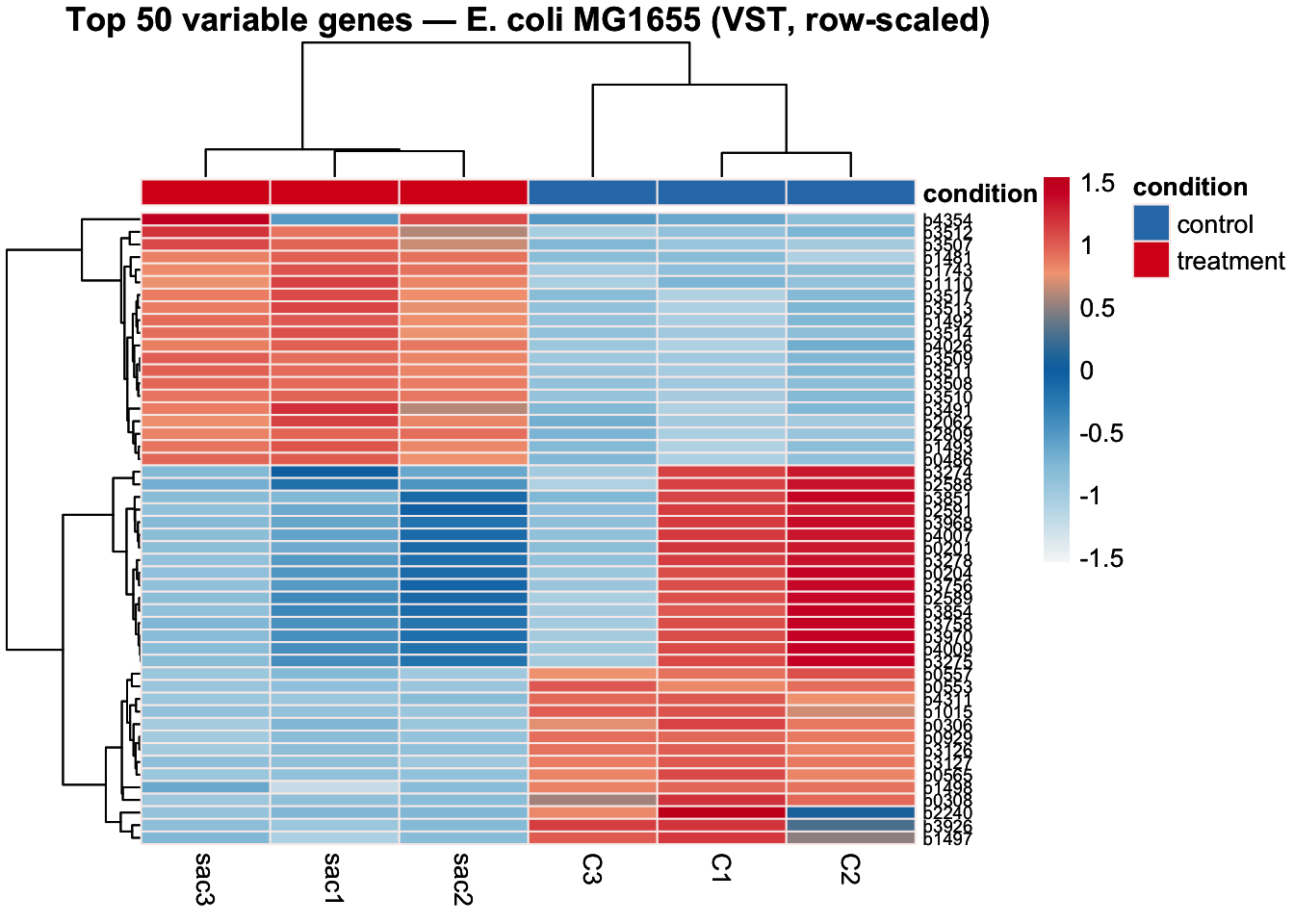
3.11 Top Variable Genes Heatmap
Clustering the 50 most variable genes across samples provides a gene-level view of the separation between conditions. Genes that are truly differentially expressed should show clear block structure, high expression in one condition, low in the other. This heatmap also helps identify whether replicates within a condition are consistent with each other.
Row-scaling (z-score per gene) is applied so that highly expressed genes do not visually dominate over lowly expressed ones.
topVarGenes <- head(order(rowVars(assay(vsd)), decreasing = TRUE), 50)
df_anno <- as.data.frame(colData(vsd)[, "condition", drop = FALSE])
anno_colors <- list(condition = cols_condition)
colsHeat <- c("#F7F7F7", "#92C5DE", "#0571B0", "#F4A582", "#CA0020")
heat_plot <- pheatmap(
assay(vsd)[topVarGenes, ],
cluster_cols = TRUE,
cluster_rows = TRUE,
scale = "row",
clustering_distance_rows = "euclidean",
clustering_distance_cols = "euclidean",
annotation_col = df_anno,
annotation_colors = anno_colors,
show_colnames = TRUE,
show_rownames = TRUE,
color = colorRampPalette(colsHeat)(255),
border_color = "#f8edeb",
fontsize_row = 7,
main = "Top 50 variable genes — E. coli MG1655 (VST, row-scaled)"
)
heat_plot